0.1.0 • Published 4 years ago
jbrowse-plugin-hgb v0.1.0
jbrowse-plugin-hgb
JBrowse2 plugin for displaying long-read alignments by hgb.
Screenshots
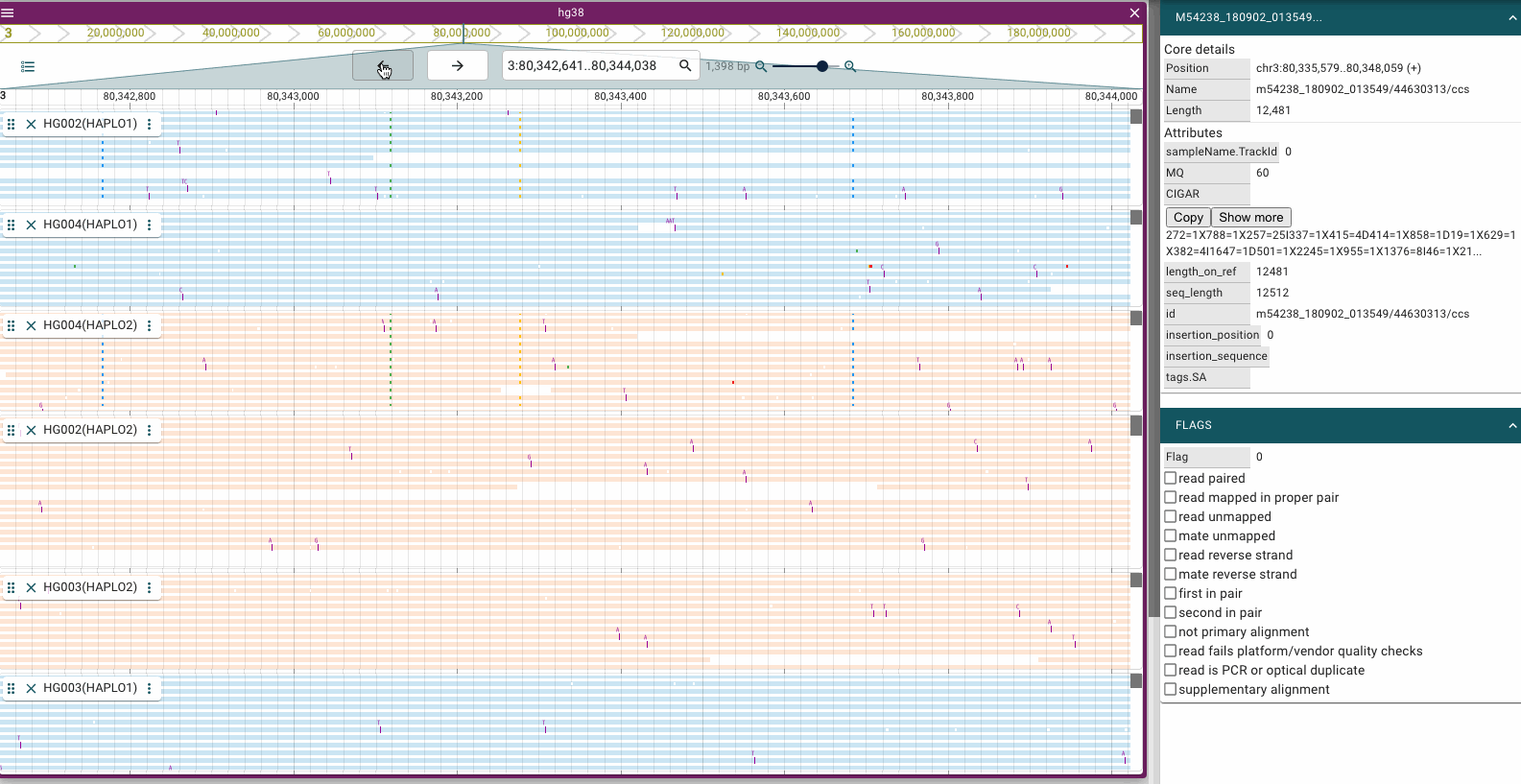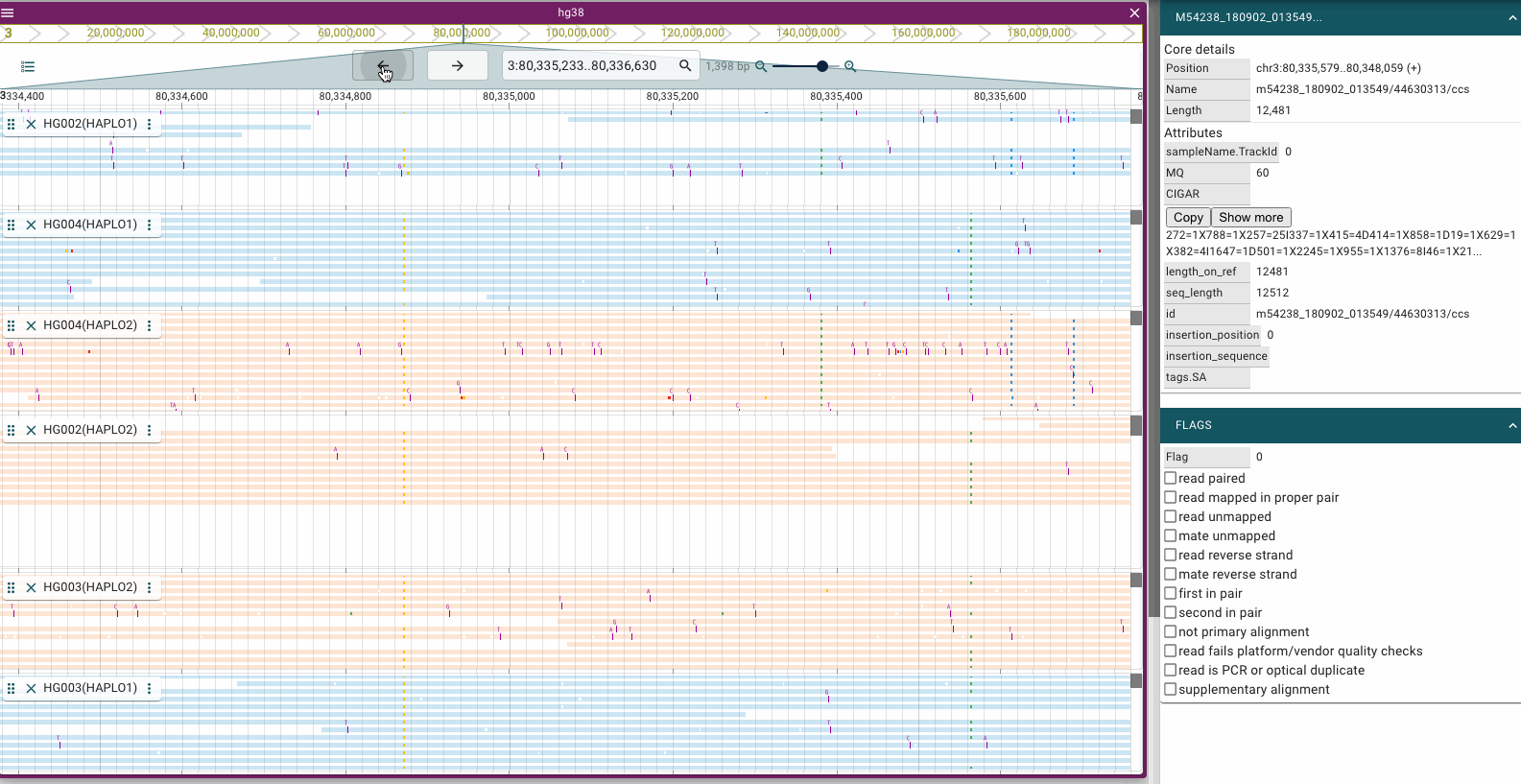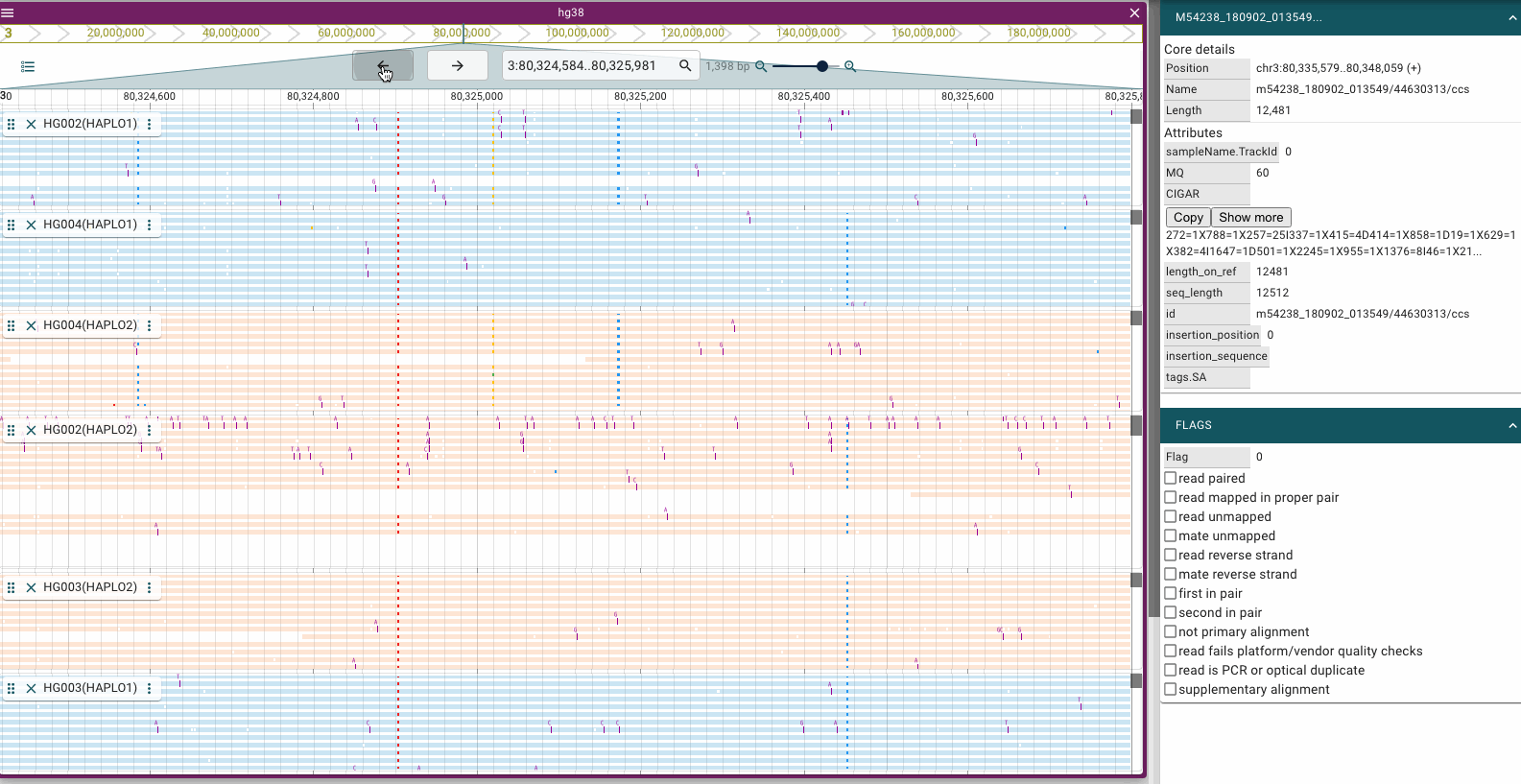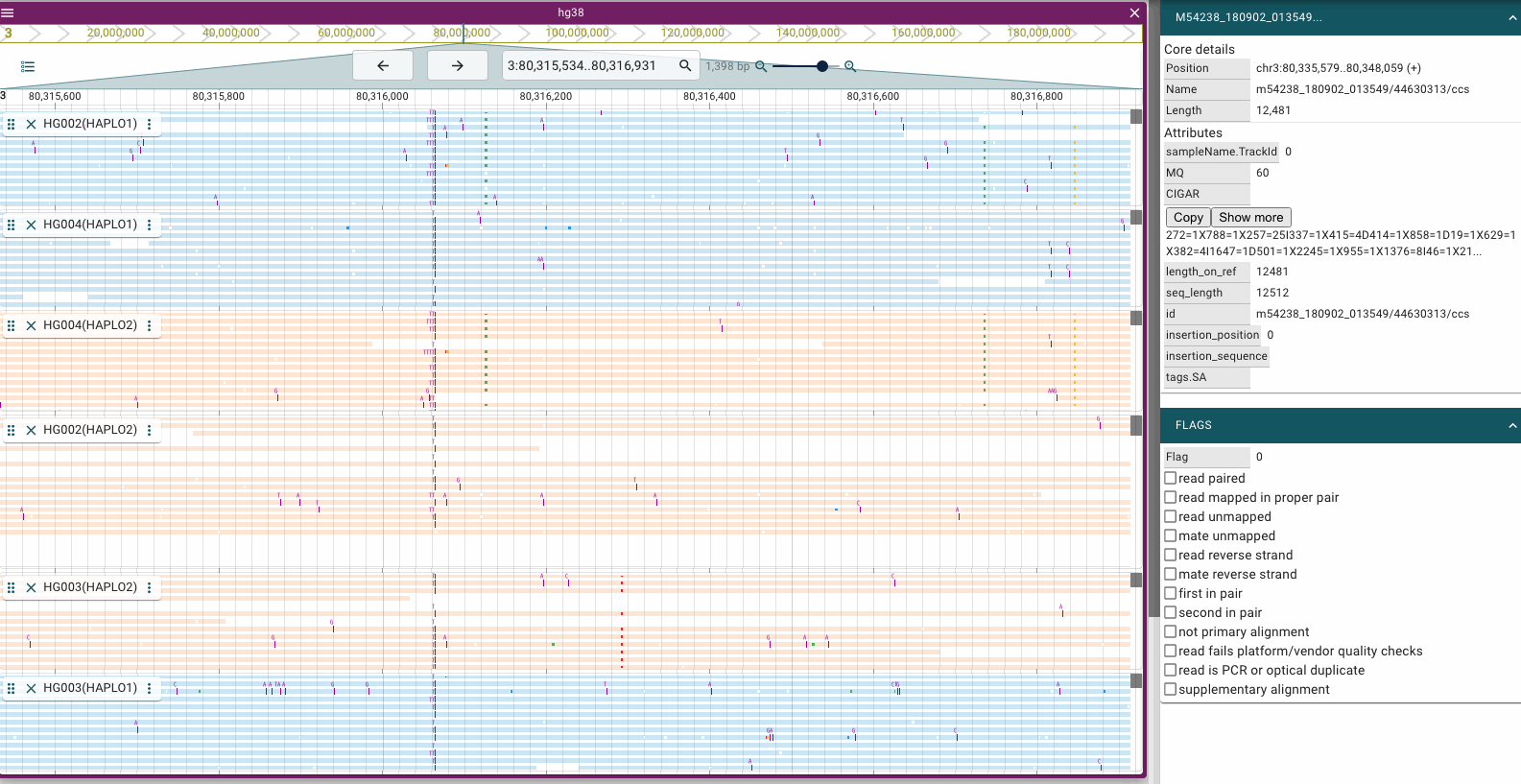
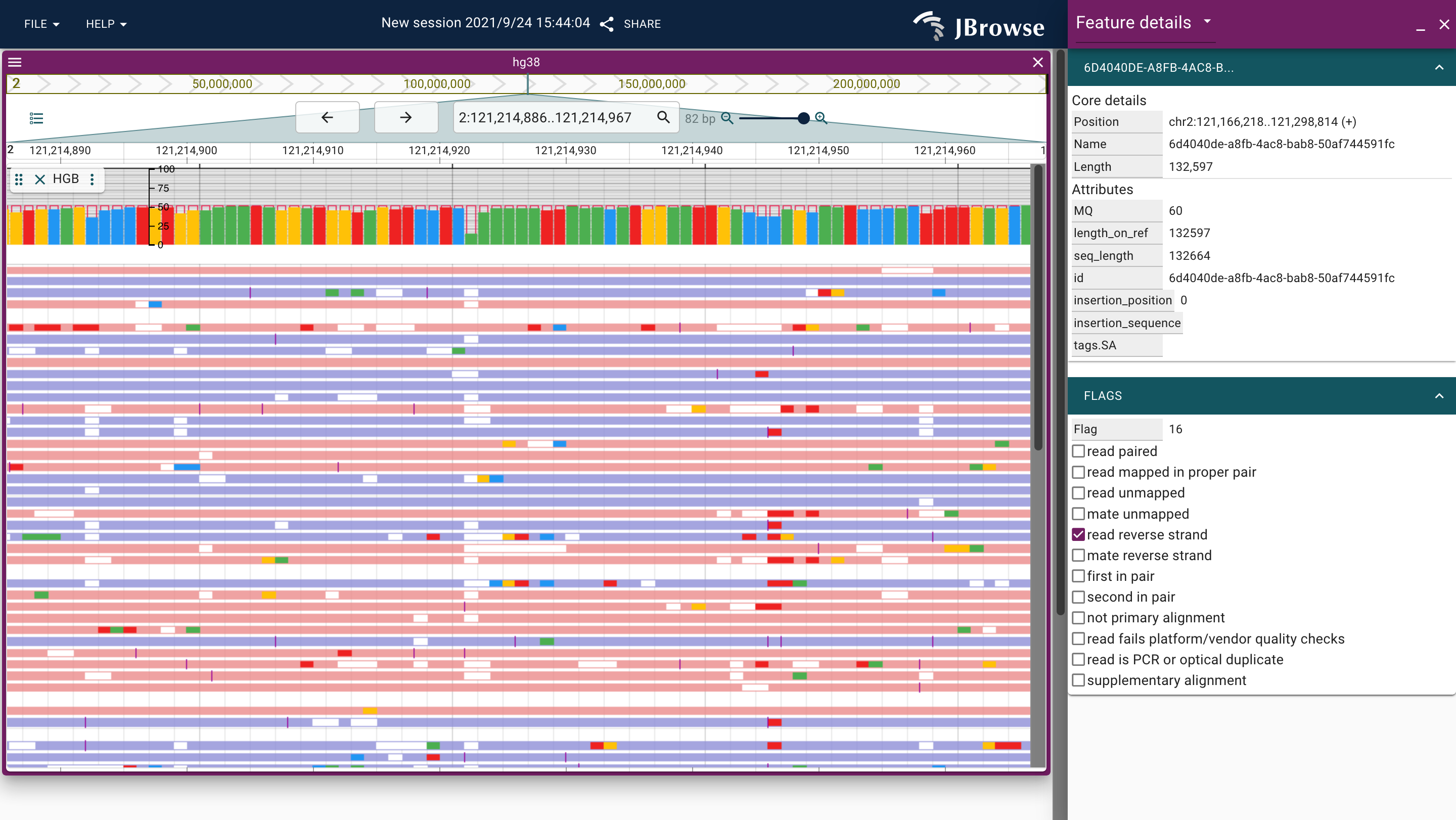
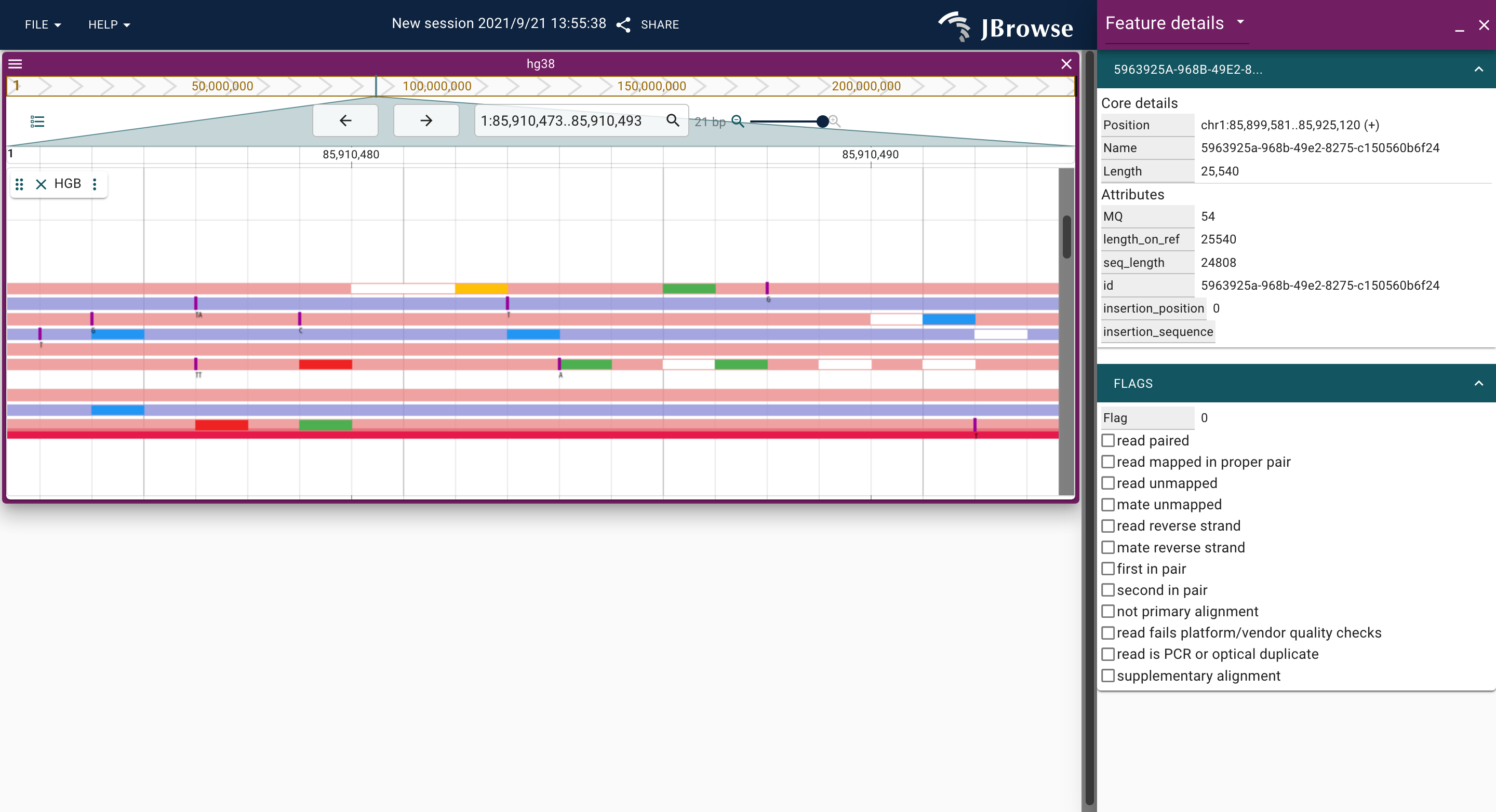
Prerequisites
- Rust nightly (cargo)
- Node.js
Getting Started
The input bam file needs to be attached MD tags by samtools calmd and indexed by samtools index.
- Docker
docker run --rm -ti -p 9000:9000 -v `pwd`:/data 6thbridge/jbrowse2-hgb:master /data/<input.bam>- Singularity
singularity build jb2-hgb.sif docker://6thbridge/jbrowse2-hgb:master
singularity -s run jb2-hgb.sif <input.bam>Access to http://localhost:9000/static/index.html.
Manual Deploy
STEP1: Serve HGB server
bash -x start_hgb_server.sh $NUM_OF_THREADS $LOCATION_OF_BAM
bash -x start_hgb_server.sh $NUM_OF_THREADS $LOCATION_OF_BAM 0.0.0.0:5000STEP2: Start JBrowse2
The following command needs to run on another shell.
bash -x start_jbrowse_hgb.sh
bash -x start_jbrowse_hgb.sh $HGB_SERVER_URL 38
bash -x start_jbrowse_hgb.sh $HGB_SERVER_URL 19You can specify reference genome as the second argument as 38 or 19 (hg38 or hg19, respectively).
Usage in jbrowse-web
Add to the "plugins" of your JBrowse Web config. The unpkg CDN should be stable, or you can download the js file to your server
{
"plugins": [
{
"name": "hgb",
"url": "https://unpkg.com/jbrowse-plugin-hgb/dist/jbrowse-plugin-hgb.umd.production.min.js"
}
]
}This plugin is currently quite basic, and there is no mouseover interactivity or drawn labels on features
For use in jbrowse/react-linear-genome-view
See DEVELOPMENT
0.1.0
4 years ago